In preparation for the launch of the NISAR mission, NASA’s Jet Propulsion Laboratory (JPL) has developed some sample NISAR products for users to explore. These sample data give users an idea of what the NISAR data products will look like once the mission is operational. The sample data provide the opportunity to get familiar with formats and develop workflows that will support NISAR datasets.
The NISAR Sample Data Product Suite page on JPL’s NISAR website provides links to all available official sample products.
ASF has developed this document to assist in working with the sample data products using various software suitable for visualizing, analyzing, and transforming geospatial data. Included are sections for working with the data in a GIS environment (ArcGIS Pro, QGIS), exploring the data using Panoply, and transforming the data using GDAL commands.
ArcGIS Visualization/Exploration

This section demonstrates how to use ArcGIS Pro to explore NISAR sample data. This documentation was generated using version 3.2.2, and assumes that you have ArcGIS Pro open, a new project open based on the Map template, and access to both the Contents and Catalog panes.
Click on a product type to view the steps for Visualization/Exploration using ArcGIS:
The sample GCOV data file is:
NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5
HDF5 Format
First, let’s explore what happens if we try to add the h5 file to the project.
- Navigate to the directory with the sample data files in the Catalog pane
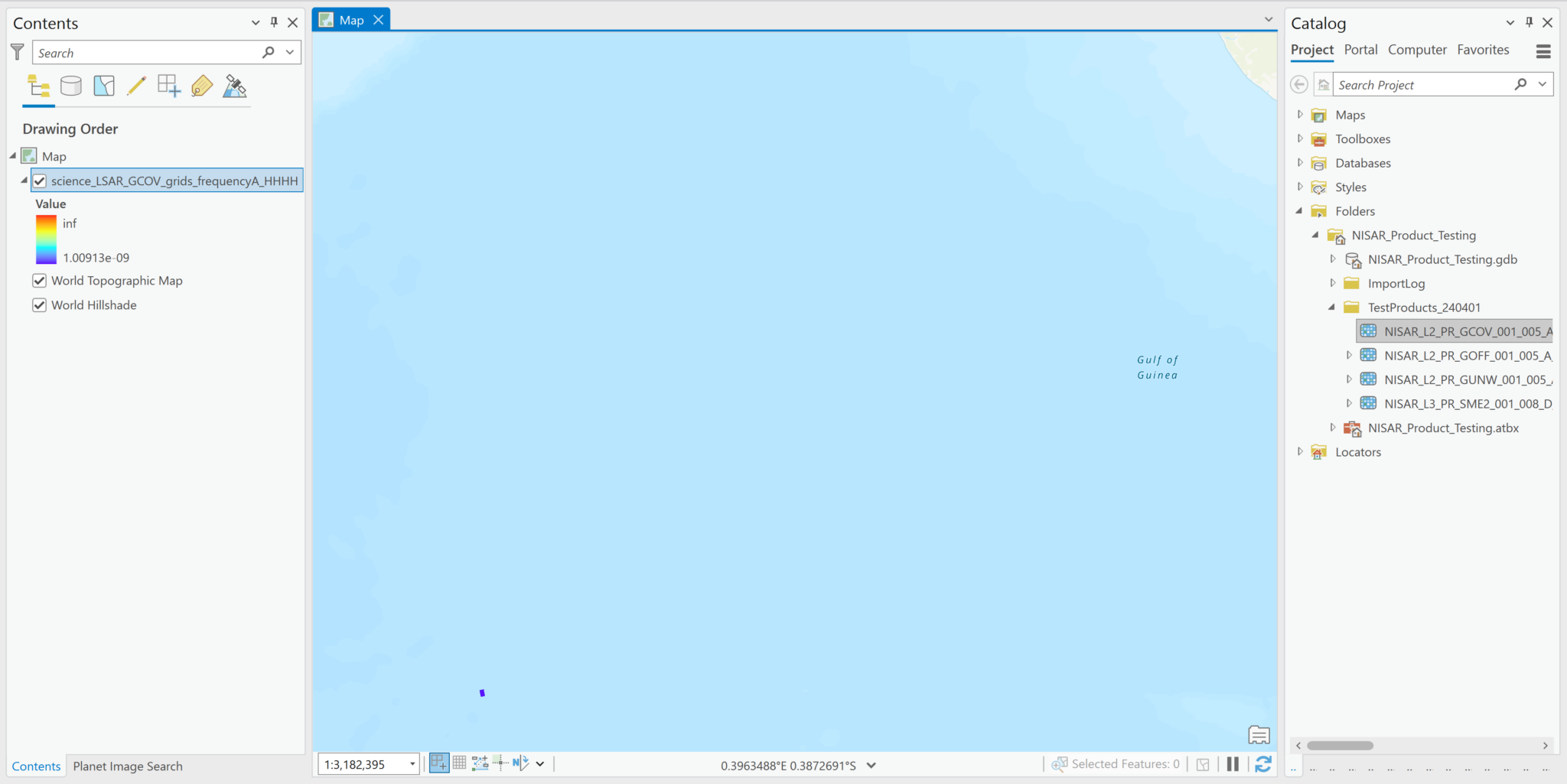
- Note that the .h5 files are listed in the Catalog pane, and are available to drag and drop into the project. If you add it to your map, it displays the L-band HH GCOV data in the map, but ArcGIS cannot access the spatial reference information, and places the raster off the west coast of Africa. In the illustration below, the raster shows up as a small purple rectangle in the ocean.
If you add the entire HDF5 file to the project, either by using the drag-and-drop method, or clicking the Add Data button and selecting the HDF5 file, you can access all of the variables in the dataset using the Multidimensional menu. Note that the Multidimensional menu only appears once you add a multidimensional dataset to your project and select it in the Contents pane.
By default, the first variable in the dataset will display in the map, but you can choose to display any of the other variables by selecting it from the drop-down list. The name displayed in the Contents pane will not change, but the map does display the data of the selected variable. You can adjust the symbology as appropriate for the variable you have selected.
Note that only the variables in the grids group will reliably display in the map. Many of the variables in the metadata group are not suitable for visualization.

- To view the raster properties, either right-click the file in the Catalog pane, or right-click the layer in the Contents pane, and select
Properties. It’s easier to see the key properties in a unified view when accessing the properties from the Catalog pane rather than the Contents pane.

- Note that the Spatial Reference is listed as
Unknown
There is no metadata formatted in a way that can be accessed using the View Metadata option (right-click file in Catalog pane), so the only insight into the dataset is what is displayed in the fields included in the dataset properties.
Treat HDF5 as NetCDF
The spatial reference information is encoded in the product using the CF metadata conventions, which were developed for use with NetCDF files. As such, if the GIS software recognizes the file as being a NetCDF, it can parse the spatial reference information and place the raster in the correct location. The current workaround for working with the NISAR HDF5 files in geospatial workflows is to rename the file with a .nc extension in place of the .h5 extension.
Change the file extension
- Browse to the HDF5 file in either a file explorer window or in the Catalog pane of your ArcGIS project.
- Edit the filename, changing the
.h5extension to.nc.- Note that once the extension has been changed, the file is no longer visible in the Catalog pane in ArcGIS. You can still add the file using the Add Data interface, but there will be no indication in the Catalog pane that this file exists.
Add Single Variable from Multidimensional Raster to Project
- Click on bottom half of the Add Data icon in the Map menu ribbon to display the menu, and select
Multidimensional Raster Layerfrom the list of options.- Note that if you just use the generic
Add Dataoption, you will be able to navigate to the .nc file, but there will be an error if you try to add it to the map, as it requires the functionality of theMultidimensional Raster Layeroption to access the variables within in the .nc file.
- Note that if you just use the generic
- Click on the folder icon and browse for the NetCDF version of the HDF5 file.
- Note that HDF5 files are indicated by a blue raster icon, while NetCDF files are indicated by a multilayer grayscale raster icon.
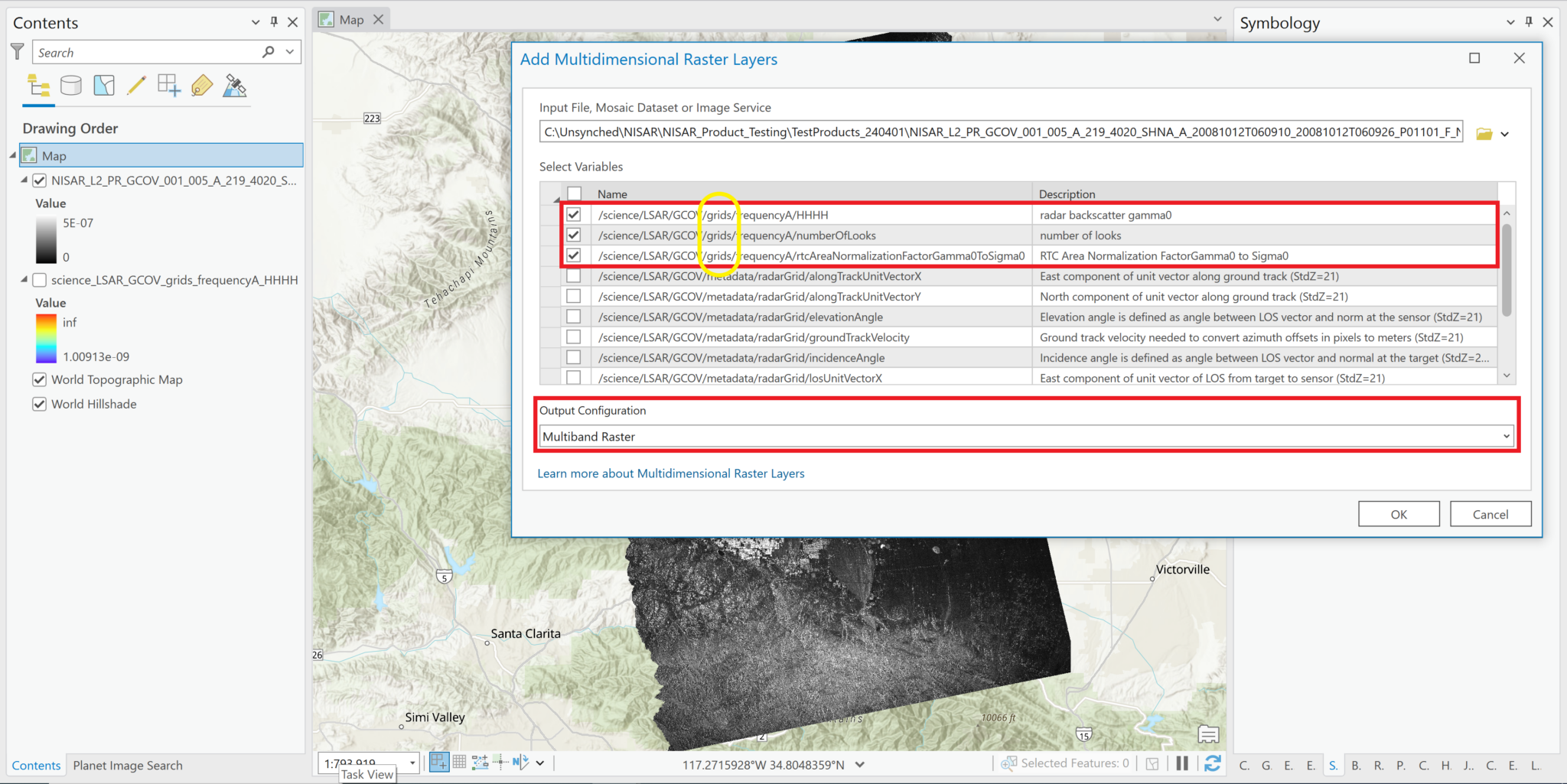
- Select the NetCDF file and click OK. The variables in the HDF5 file will be listed.

- Check the box next to the radar backscatter variable to add it to the project. In this example, only one polarization is available.
- Use the drop-down menu for the Output Configuration parameter to select
Multiband Raster.- It is critical to select this option; ArcGIS will not render the data if any of the multidimensional options are selected for the Output Configuration.
- It is possible to select multiple variables to combine into a single multiband raster, if desired. For most geospatial analysis applications, this is probably less convenient than adding the variables separately, but it is a convenient way to add many variables to the map at once for a quick exploration. Refer to the Add Multiple Variables from Multidimensional Raster to Project section for more details.
- Click OK to load the variable into the map.

Adjust Stretch
As with most SAR backscatter images in power scale, the image appears mostly black when it’s added to the map. You can adjust the stretch settings to view features in the image.
- Right-click on the layer in the Contents pane and select Symbology
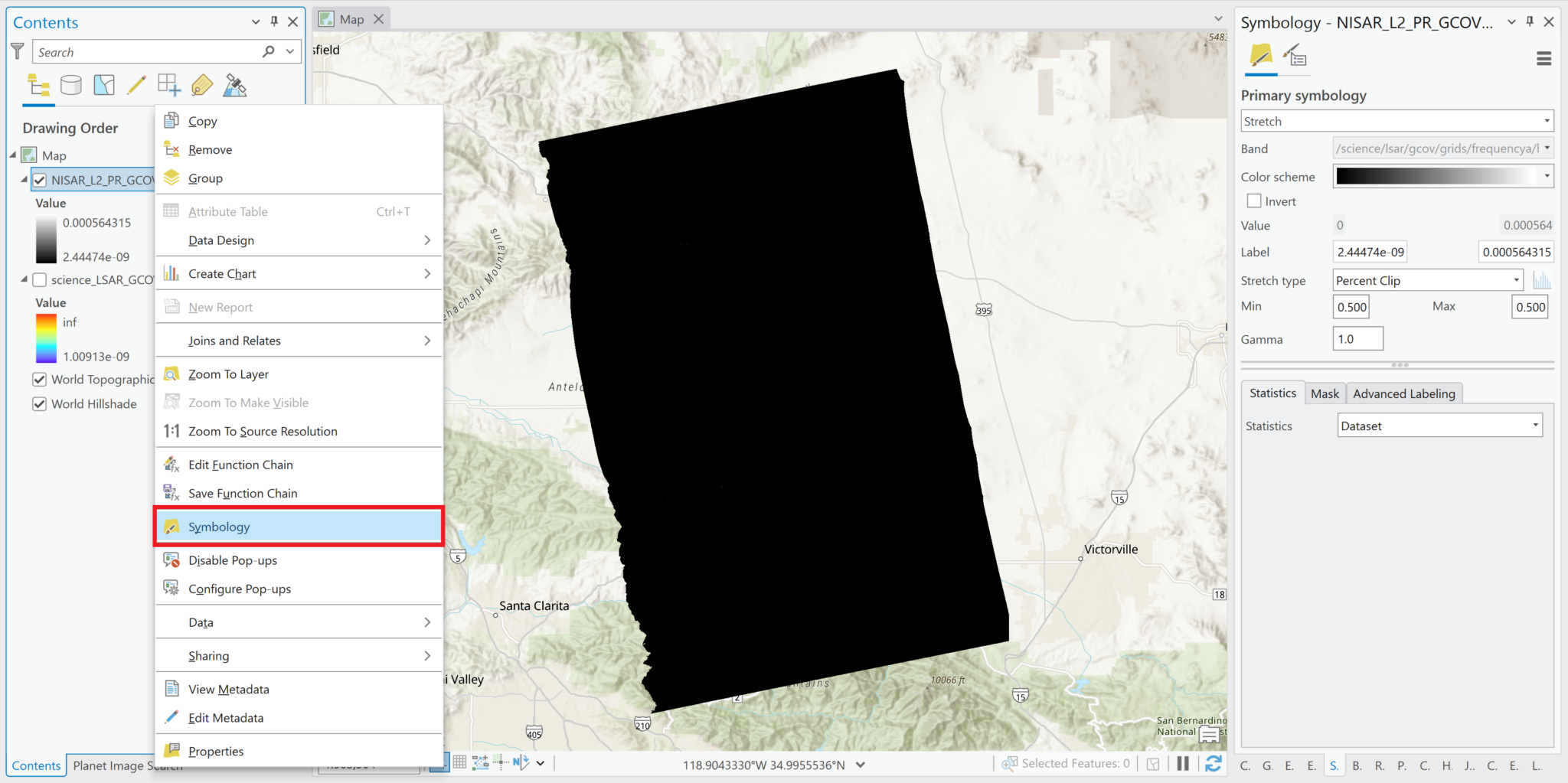
- Adjust the settings to visualize the layer as desired. Common approaches include:
- Set the Stretch Type to
Standard Deviationand adjust the Number of standard deviations value as desired. – For this sample image, 0.5 seems to be a reasonable value. – Changing the Gamma value from 1 to 2 reduces the amount of contrast in the mid-level grayscale values, adding more nuance to the image.

- Set the Stretch Type to
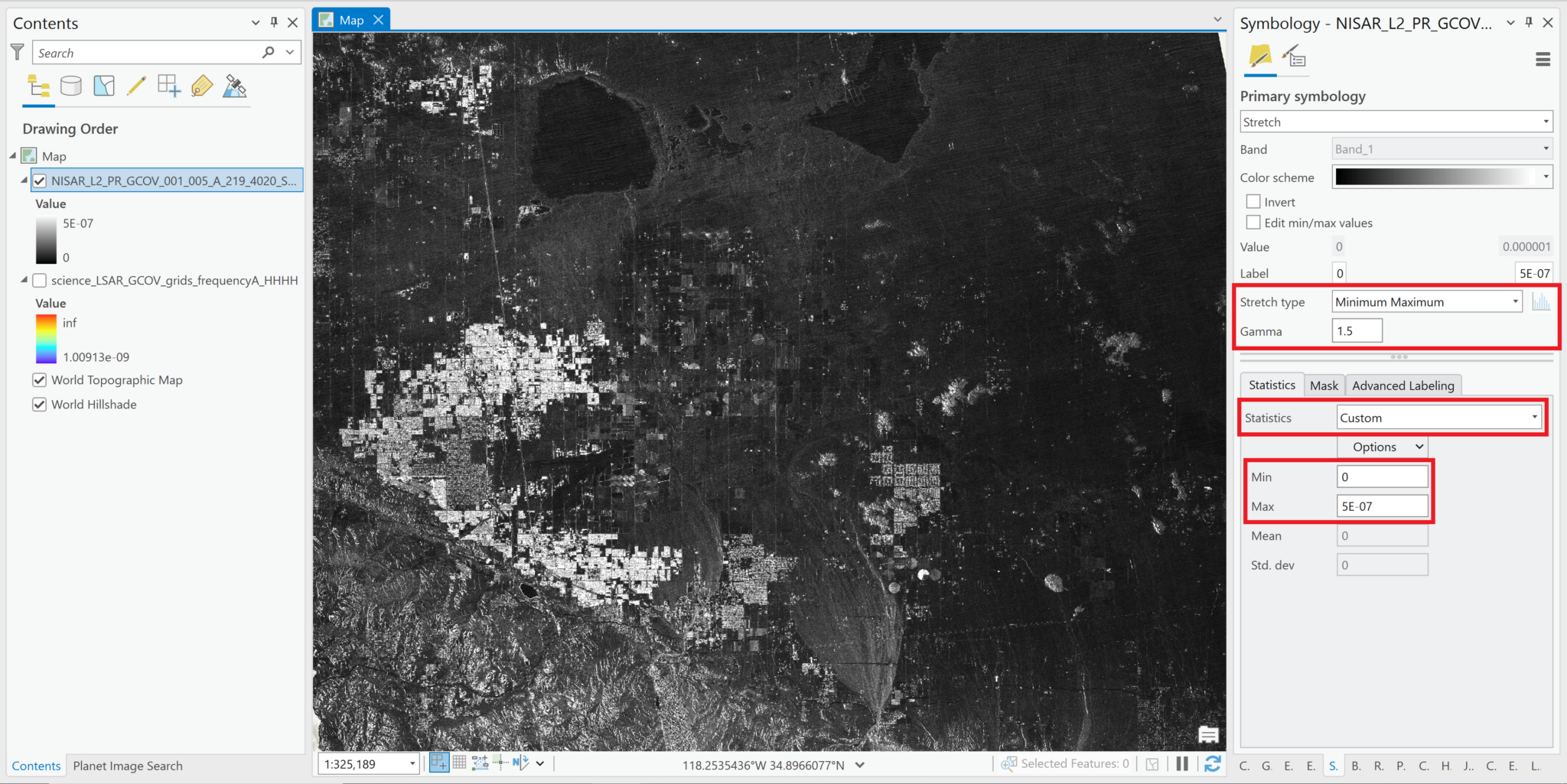
Minimum Maximumand selectCustomfrom the Statistics drop-down menu. This allows you to set a specific range of values to use for the grayscale in the Min and Max fields. Use the tab button after entering each value so that the label field is updated appropriately.- Normally, we would expect the range power values for RTC products to fall between about 0 and 0.03, but the values in this sample image are much closer to zero. It appears that the values are the covariance measurements, rather than backscatter normalized to gamma-0 power. Most values fall between 0 and 0.0000005.
- Again, increasing the Gamma setting from 1 can also improve the visualization.

Add Multiple Variables from Multidimensional Raster to Project
This option could be particularly useful when there are multiple polarizations that you want to visualize quickly, especially if you want to use the different polarizations in different channels to display an RGB Decomposition.
- Follow the same steps as in Add Single Variable from Multidimensional Raster to Project to select the NetCDF and list the variables.
- Check the boxes next to the variables you want to include.
- Only the variables in the
/grids/group in the HDF5 file are suitable to use when adding multiband rasters with multiple variables.
- Only the variables in the
- Select the Multiband Raster from the Output Configuration drop-down menu, and click OK to add the multiband raster to the image.
Adjust the symbology to visualize the different bands as desired. By default, if three different variables are included, the image will be visualized as an RGB image with each of the variables assigned to a different band. If fewer than three variables are included, the image will be a grayscale stretch, and you can select which variable to display from the Symbology menu.
It is critical to allow the imagery to render fully each time you change a symbology setting before adjusting another symbology setting.
For example, if you change to a Standard Deviation stretch, allow the image to render fully before changing the number of standard deviations.
Here are some notes on visualization options:
- In some cases, it may make sense to keep the RGB visualization. This could be particularly powerful if you’re interested in combining the variables to display particular properties. Unfortunately, the best use case is probably for images with multiple polarizations, but the sample data only includes HH.
- In this illustration, the three variables available in the grid group are assigned to the RGB channels. Because the backscatter values are so small compared to the range of values in the other two variables, there is essentially no red in the image.
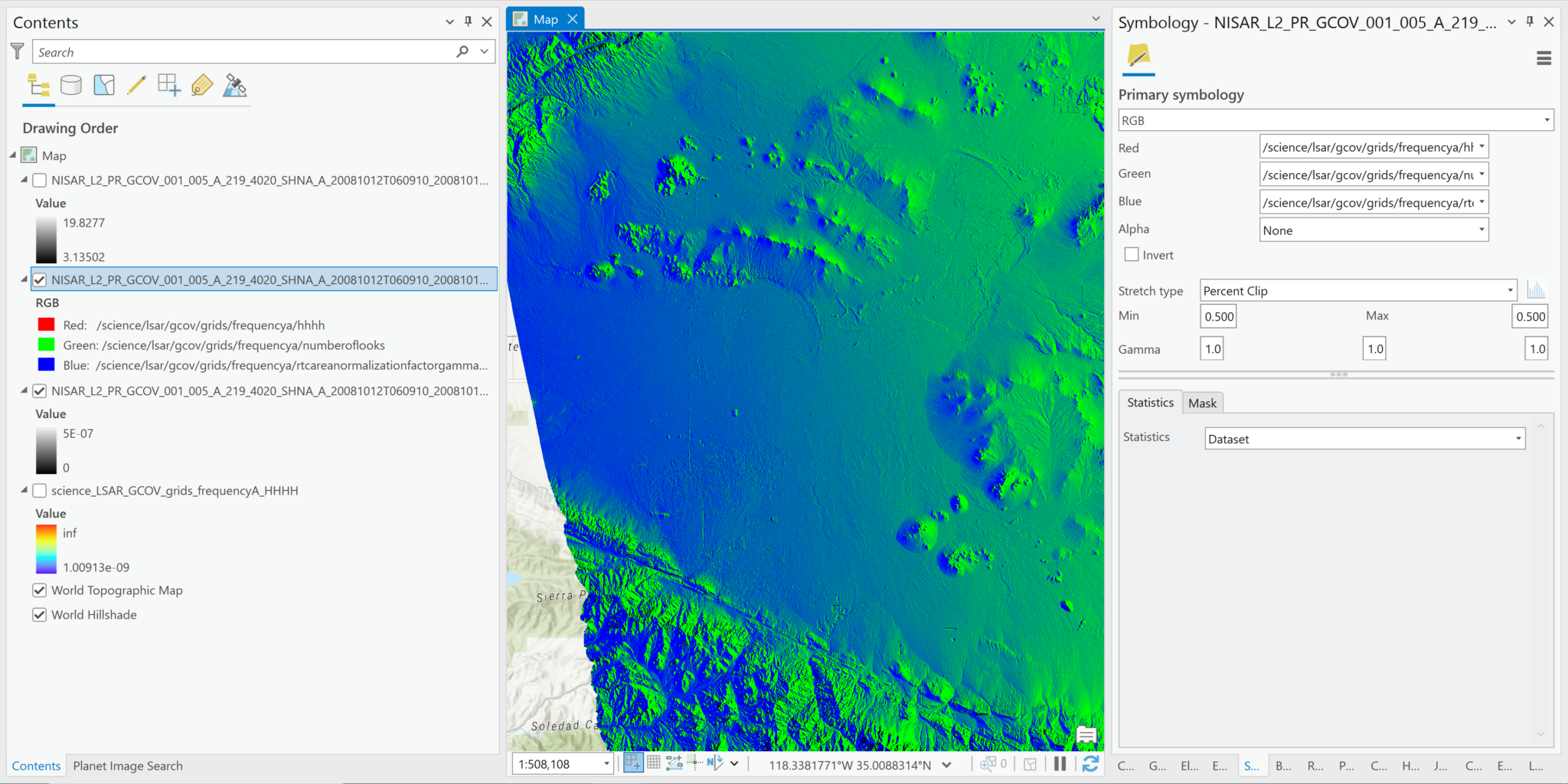
- You can also adjust the stretch type within the RGB image to highlight particular values/features, although the variables available for use in this example do not offer a particularly compelling use case for this functionality.
- In this illustration, changing the stretch type to Standard Deviation allows you to see areas with high backscatter in relation to the other two variables.
- Setting the number of standard deviations to 0.5 would make the range of backscatter values even more obvious.

- You can change the RGB to a Stretch visualization, where you choose to visualize one variable at a time. If you want to make this change, it is best to do it before making any symbology adjustments to the RGB settings.
- If you change from RGB to Stretch right away, the variable names are displayed in the list of bands, making it easy to select the variable you want to display. Once there have been edits and changes, especially if you change the bands assigned to the different RGB channels, you can lose that information, and it just lists “Band_1, Band_2, Band_3” rather than the variable names.
- Use the Band drop-down menu to select the variable to display.

You can try to adjust the stretch characteristics as desired for the selected variable, but the connection to the data source can sometimes be lost, especially if ArcGIS needs dataset statistics in order to apply the stretch settings. For fine-tuning grayscale visualizations, it’s better to add variables individually rather than stacking them into a multiband raster.
It is also possible to change from a Stretch symbology to an RGB symbology.
- This would be particularly useful for dual-pol imagery. When you add two variables, it displays as a stretch grayscale image by default.
- Change the symbology from Stretch to RGB, and select the band to use for each color channel. A common approach for dual-pol imagery is to apply the co-pol (usually HH for NISAR) data to the Red and Blue channels, and the cross-pol (usually HV for NISAR) data to the Green channel.
The sample GUNW data file is:
NISAR_L2_PR_GUNW_001_005_A_219_220_4020_SH_20081012T060910_20081012T060926_20081127T060959_20081127T061015_P01101_M_F_J_001.h5
Note that most of the variables in this file have float or integer pixel types, but the wrapped phase variable has a complex pixel type. ArcGIS can display all of the variables, but refer to the Wrapped Phase section for details on how to appropriately visualize complex datasets. This table describes the pixel types for all of the variables included in the grids group of the h5 file:
| Variable | Pixel Type |
|---|---|
| /science/LSAR/GUNW/grids/frequencyA/pixelOffsets/HH/alongTrackOffset | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/pixelOffsets/HH/correlationSurfacePeak | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/pixelOffsets/HH/slantRangeOffset | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/unwrappedInterferogram/HH/coherenceMagnitude | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/unwrappedInterferogram/HH/connectedComponents | 32-bit unsigned integer |
| /science/LSAR/GUNW/grids/frequencyA/unwrappedInterferogram/HH/ionospherePhaseScreen | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/unwrappedInterferogram/HH/ionospherePhaseScreenUncertainty | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/unwrappedInterferogram/HH/unwrappedPhase | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/unwrappedInterferogram/mask | 8-bit char |
| /science/LSAR/GUNW/grids/frequencyA/wrappedInterferogram/HH/coherenceMagnitude | 32-bit float |
| /science/LSAR/GUNW/grids/frequencyA/wrappedInterferogram/HH/wrappedInterferogram | 64-bit complex |
Data from this file can be accessed using the same workflow described for the GCOV Product.
- Change the file extension from .h5 to .nc
- Use the Multidimensional Raster Layer option in the Add Data menu to browse to the .nc product and select the variables to add to the map.

The most common variables to visualize are the wrapped and unwrapped interferograms and the coherence magnitude, but any of the variables in the \grids\ subgroup are rasters that can be displayed in a GIS environment.

As with the GCOV products, the output configuration selected when adding variables must be Multiband Raster, but you can choose to add individual variables as separate rasters (technically a single-band raster, but still defined as a multiband raster), or select multiple variables to be added to the map together in a single multiband raster (that truly includes multiple bands, each with a different variable).
In general, if you’re interested in optimizing the visualization of a raster, it’s better to add the variable of interest by itself, rather than as a multiband raster that includes other variables. The more you tweak the symbology, the more likely you are to lose the connection to the file, and have to start over. If there’s only one variable included in the multiband raster, it seems to be more robust to symbology adjustments. That said, the connection to the source data can be lost even with a single-variable raster, especially if a symbology adjustment is made that calls for statistics.
When you first add a raster to the map, the variable names usually show up in the list of bands in the Symbology tab. This is the case both for rasters with multiple variables included, and those with just a single variable displayed in the raster, both for greyscale and RGB. This is an easy way to determine which variable you’re exploring.
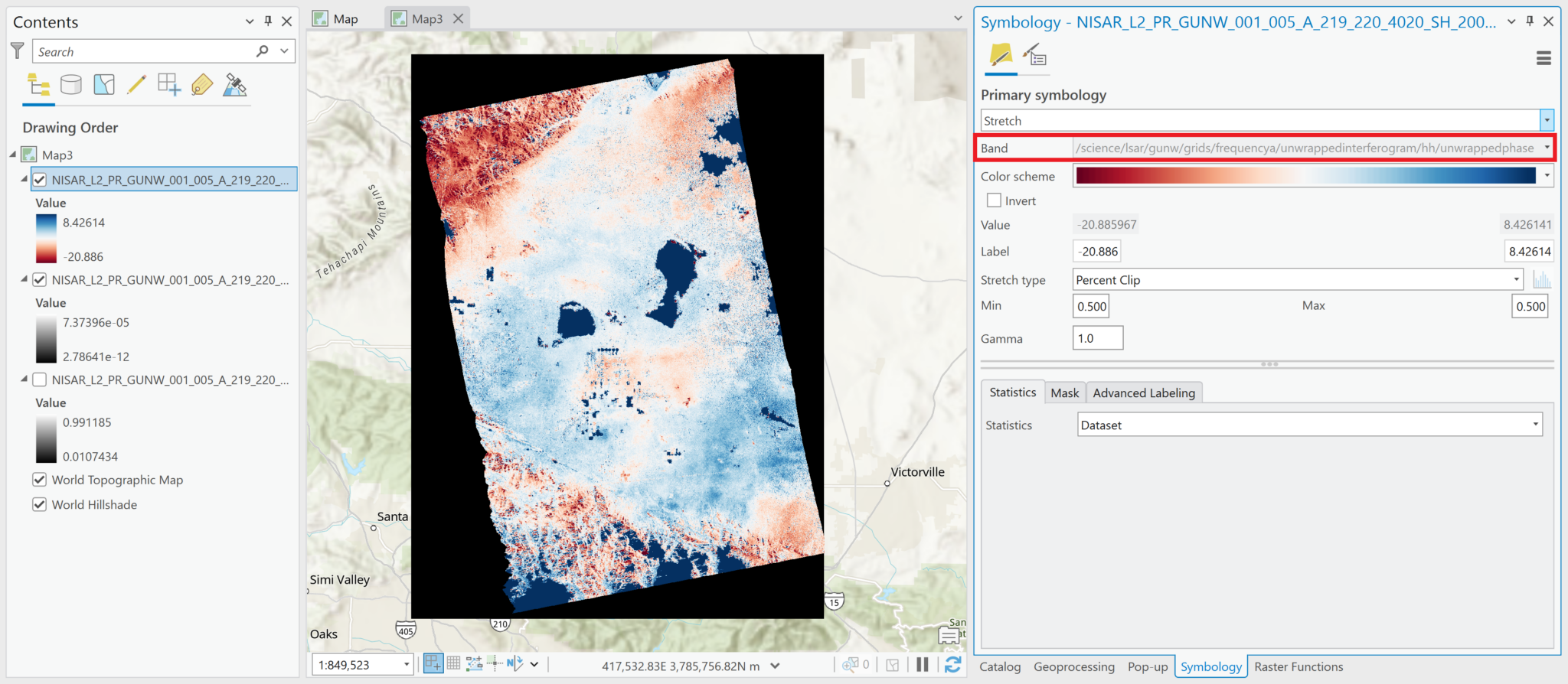
Sometimes the band name gets lost, however, especially after changes are made to the symbology. Instead of the variable name being displayed in the Symbology tab, there is just the generic Band number listed. Because the name of the variable is not displayed in the contents pane by default, it is easy to lose track of which variable the raster represents. You may want to rename the raster layer in the Contents pane, or add an identifier to the end of the existing layer name.
In the illustration below, the wrapped interferogram variable name is no longer displayed in the Band field of the Symbology tab, so we can identify it by right-clicking the layer in the Contents pane, and selecting Properties. In the Source section, the Name field displays both the filename and the variable path.
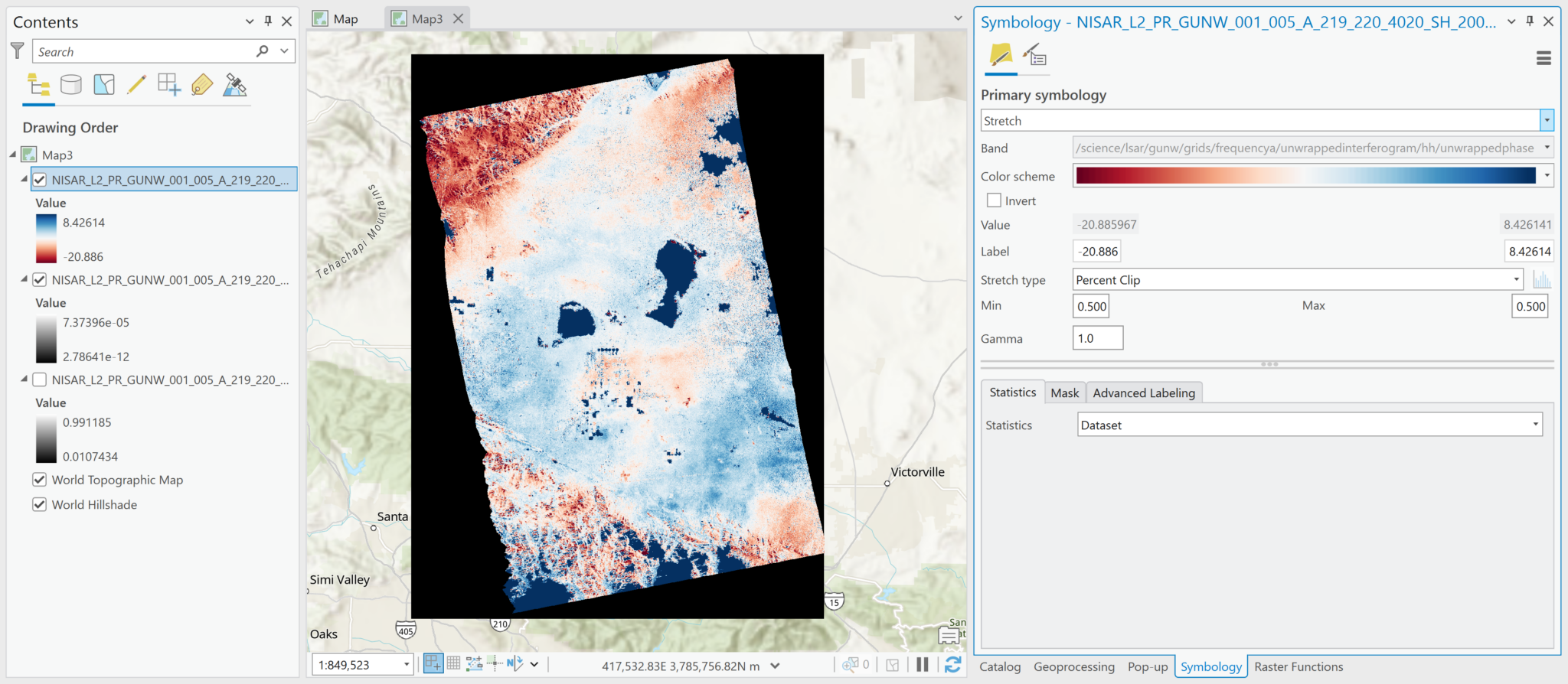
Note that there is an option to Set Data Source in the properties. Normally, if you lose a connection to a dataset, you can use that option to select the source file and repair the connection. Unfortunately, because the .nc files are not recognized as ‘files’ by ArcGIS, this is not a viable option to use if you lose the connection to the file while adjusting the symbology. If you try to navigate to the folder with the .nc files, only the .h5 files are listed.
Wrapped Phase
The wrapped phase variable has a complex pixel type, containing both real and imaginary components that can be used to calculate either the amplitude or the phase. When looking at an interferogram to detect and quantify surface deformation, we’re interested in differences in phase rather than in amplitude. By default, however, ArcGIS displays the image in terms of the amplitude values instead of the phase, with pixel values ranging from 2.78641e-12 to 7.37396e-05 for the sample dataset.
Starting with version 3.0, ArcGIS Pro provides a ‘Complex’ raster function, which can be used to display the phase values instead of amplitude values. It is included in the SAR section of the raster functions panel in ArcGIS Pro.
- Add the wrapped interferogram variable to the map
- You may want to rename the layer to ‘wrapped interferogram’ if there are already multiple layers from the NetCDF file, as it will make it easier to select the appropriate layer in the raster function interface
- From the
Imagerymenu, selectRaster Functionsto open the Raster Functions panel - Type
Complexin the search bar at the top of the Raster Functional panel, and click on theComplexraster function - Select the wrapped interferogram layer in both the Raster and the Imaginary Raster fields
- Select
Phasefrom the Value Type drop-down menu - Click the
Create New Layerbutton to display a layer showing the Phase values instead of the Amplitude values
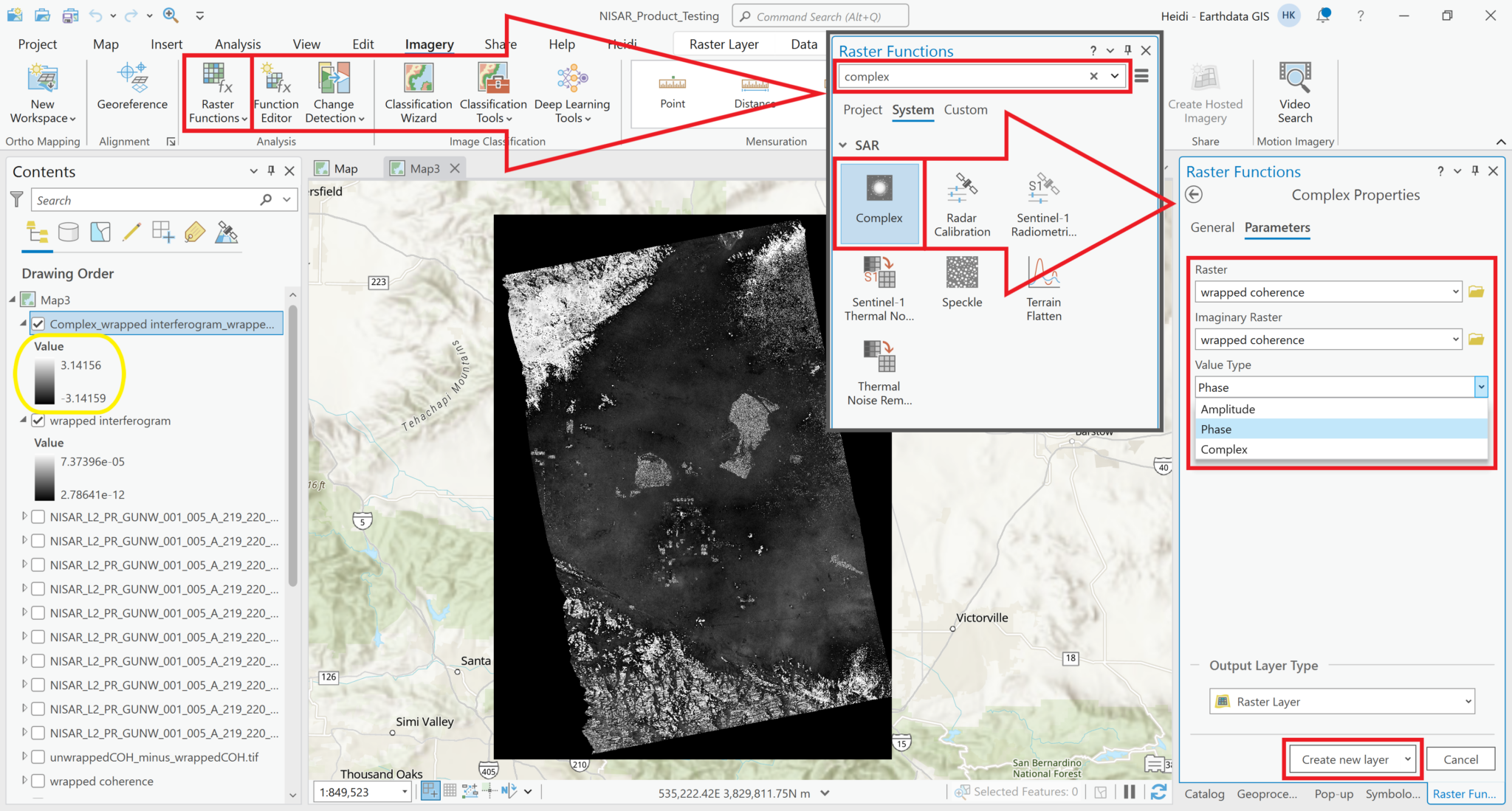
The values displayed in the image now range from -pi to pi, which is what we would expect for a wrapped interferogram.
Note that the phase layer is a virtual raster that is displayed on-the-fly in your project, but you can export the layer as a stand-alone raster if you would like to save it for future use in other applications.
Coherence
There are two different coherence magnitude variables included: unwrapped coherence magnitude and wrapped coherence magnitude. They have slight differences in their values, but it’s unclear why. There doesn’t appear to be any guidance on which one would be preferable to use for particular applications.
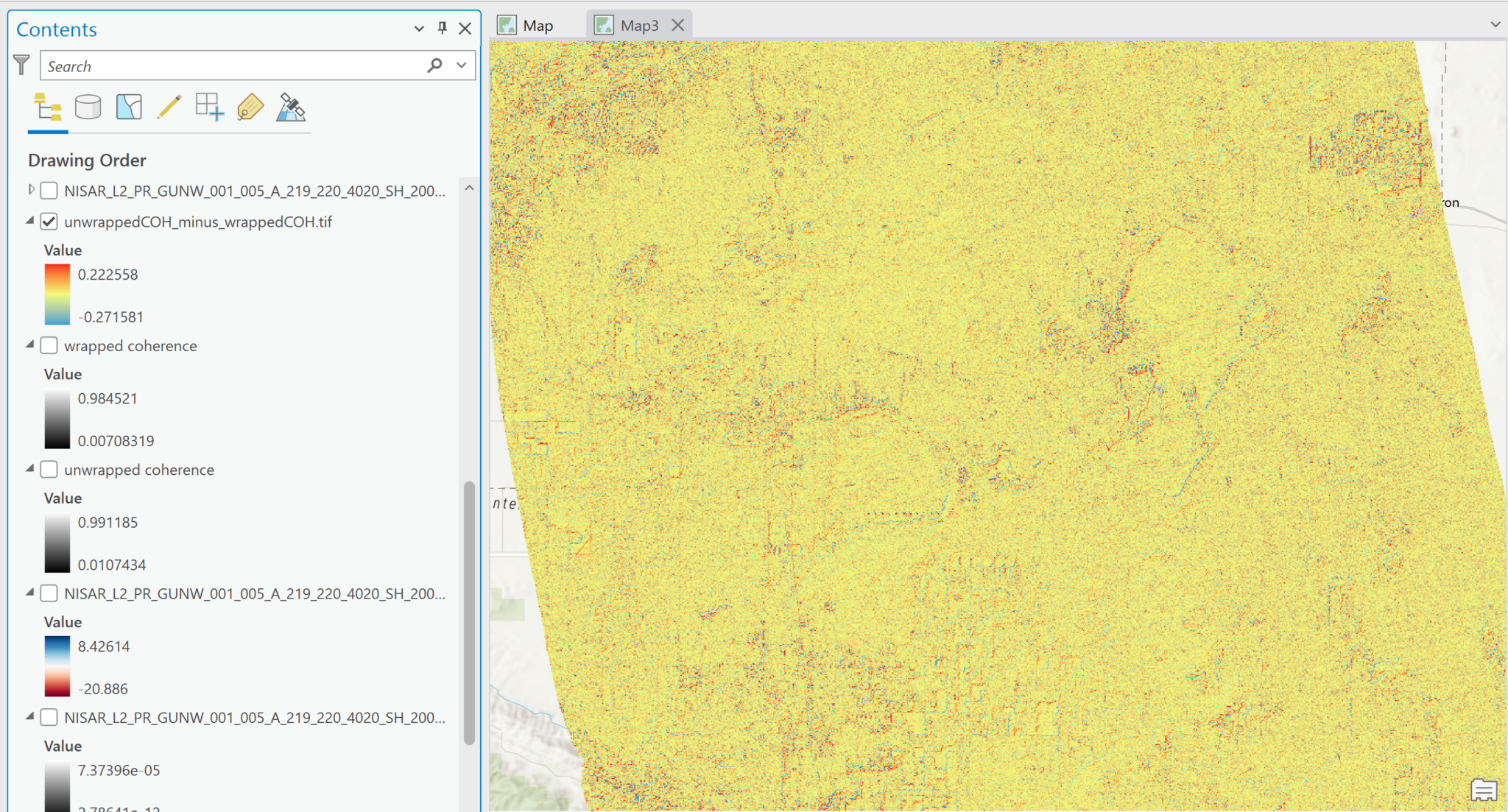
Both versions have values ranging from 0 to 1, as expected for coherence values, and are easily displayed in grayscale without having to adjust the scale. In this illustration, both images were set to have a Min-Max stretch from 0 to 1 so that they could be compared visually, but the actual dataset min-max values are also displayed next to the functional scalebar for each image.

Mask
There is a layover-shadow mask included as a variable. It uses various integers to represent certain conditions, but I have not been able to find a reference for what the values represent. It is most useful to display this layer with a Discrete symbology, which applies a different color for each pixel value. In the sample data, there are three different pixel values present: 0, 2 and 127.
0 appears to indicate that there are no quality flags, and 127 indicates the padding pixels around the valid data. The pixels with a value of 2 are generally in the mountains, but it’s unclear if it indicates layover, shadow, foreshortening, or a combination of multiple conditions.

Click on the color patch next to each value in the symbology pane to manually select colors for each individual value. It can be helpful to set the 0 values to be transparent when viewing the mask in relation to an interferogram raster.
The sample GOFF data file is:
NISAR_L2_PR_GOFF_001_005_A_219_220_4020_SH_20081012T060910_20081012T060926_20081127T060959_20081127T061015_P01101_M_F_J_001.h5
There are 21 variables included in the grid group, seven variables for each of three layers. They are all 32-bit float datasets, but all of the pixel values for all of the layers in the sample dataset are 0, so there is nothing to visualize.
The sample SME2 data file is:
NISAR_L3_PR_SME2_001_008_D_070_4000_QPNA_A_20190829T180759_20190829T180809_P01101_M_P_J_001.h5
There are three different variables containing soil moisture data: DSG (Disaggregated), TSR (Time Series Rates), and PMI (Physical Model Inversion).
Note that the spatial reference for the SME2 product is defined as WGS84 Geographic Coordinate System (EPSG 4326), but it displays in an unexpected way when added to the map.
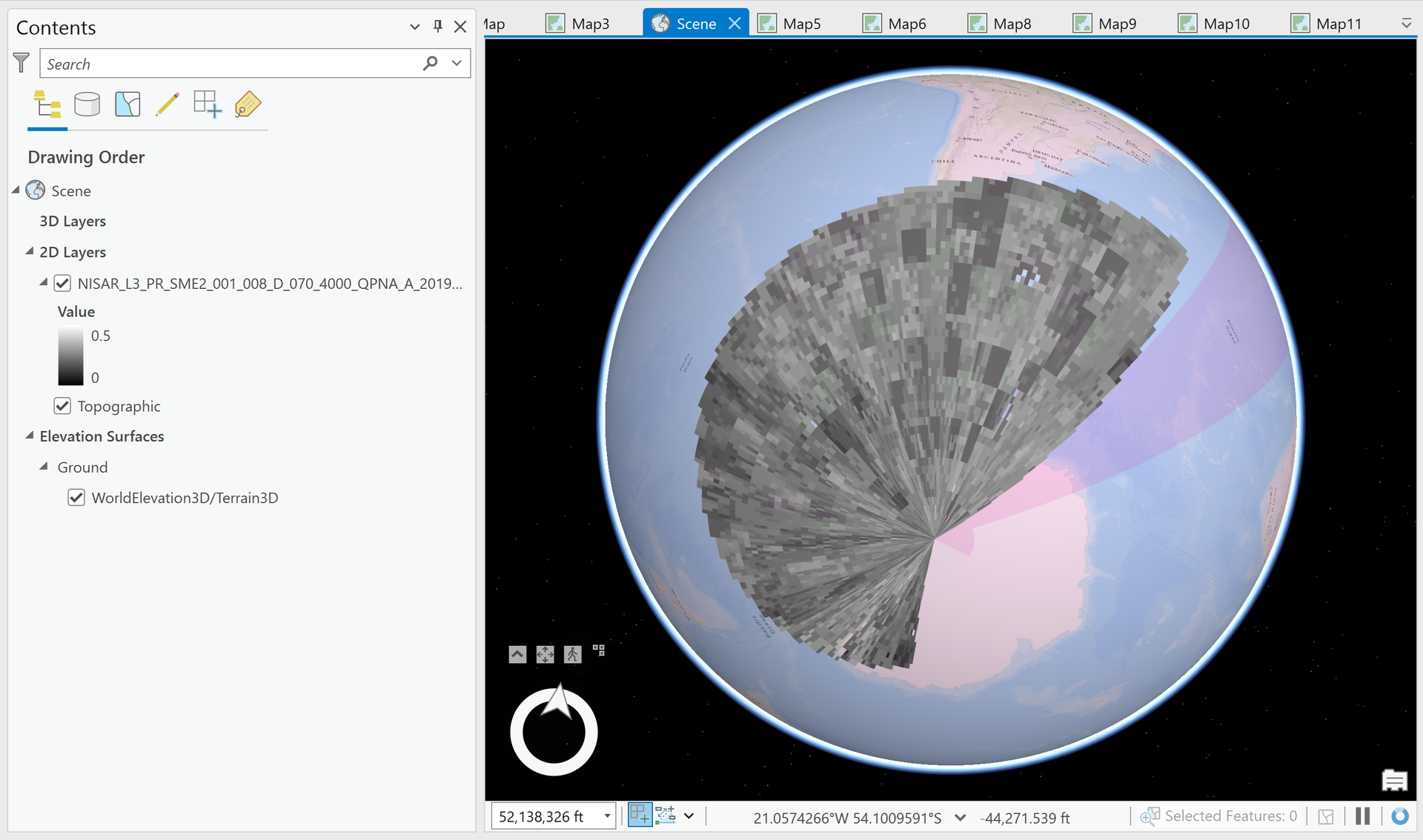
It is unclear how to work with this dataset to project it to the correct location on the earth’s surface.
Note that pixels with no data are designated with a pixel value of -9999. Even if the raster could be projected correctly, the visualization would need to be adjusted to accommodate these values.

To adjust the visualization to exclude the -9999 values from the grayscale and treat them as background pixels:
- Right-click on the soil moisture layer, and select
Symbology - Click the
Masktab, check the box toDisplay background value - By default, the background is transparent
- If you want to see the extent of the padding pixels, you can select a color (the illustration used pink)
- Once a color has been selected, click on the color swatch again and select ‘Color Properties’, where you can apply a transparency value (the illustration used 90%)

- Change the
Stretch typeto Minimum-Maximum - Select the
Statisticstab - Use the drop-down menu to set the Statistics field to
Custom - Change the value in the Minimum field from
-9999to0and press the tab button to update the scalebar label

QGIS Visualization/Exploration

This section demonstrates how to use QGIS to explore NISAR sample data. Documentation was developed using version 3.36, and only the Windows operating system allows users to interact with all the datasets without crashing the software. Refer to the QGIS Crash Issue section below for more information.
Click on a product type to view the steps for Visualization/Exploration using QGIS:
In order to load the images into QGIS, you will have to replace the .h5 file extension with .nc. This is because the NISAR data is actually formatted like a NetCDF file, rather than HDF5. If you try to use the regular .h5 files, QGIS will likely crash (with no error or warning) when you choose a layer (refer to the QGIS Crashes section for more information). If you load the .h5 file, QGIS may continue to crash even after renaming to .nc. This may be fixed by deleting the .aux.xml file that QGIS creates in the same directory as the dataset.
Once you have the file named, you can simply drag and drop it into the QGIS window. This should bring up a prompt to select the desired layers. In our case, we want the layer that ends with HHHH.

You should now have something that looks like this:

By default, QGIS scales the color map based off of the min and max. This leads to a dark image. We can fix this by changing the way that QGIS scales the color map. We can do this by right-clicking the layer and selecting Properties. We can then go to Symbology -> Min / Max Value Settings and select Cumulative Cut Count.

Now, things should look much better.

If you would like color, rather than a black-and-white image, you can click on the Render Type drop-down box in the Symbology page and select singleband psuedocolor. For this, you will also want to use Cumulative Cut Count.
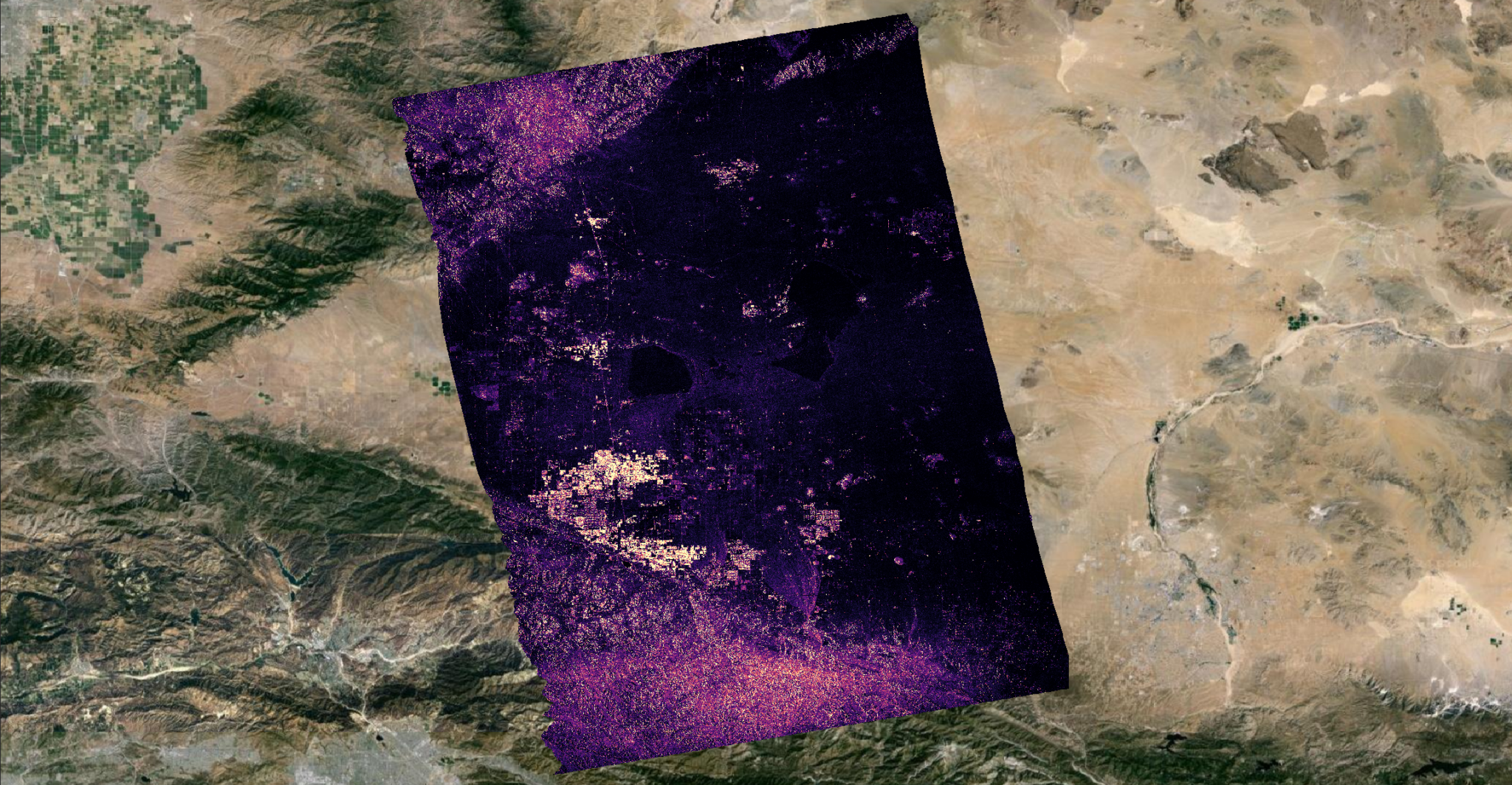

If you’re experiencing instant-crashes, even if you’ve changed the file extension to .nc, refer to the QGIS Crashes section.
Selecting the Desired Groups/Layers/Sub-Datasets?
Just like the GCOV products, we will first need to replace the .h5 extension with .nc, otherwise the projection information will be incorrect. Once the extension is replaced, we can drag the file into QGIS as normal and then select the desired layers. For this demonstration we will select the unwrappedInterferogram/HH/coherenceMagnitude and unwrappedInterferogram/HH/unwrappedPhase layers.

Unwrapped Phase
Now that the layers are added, we can right-click and zoom to the unwrapped phase layer. You should have something that looks like this:
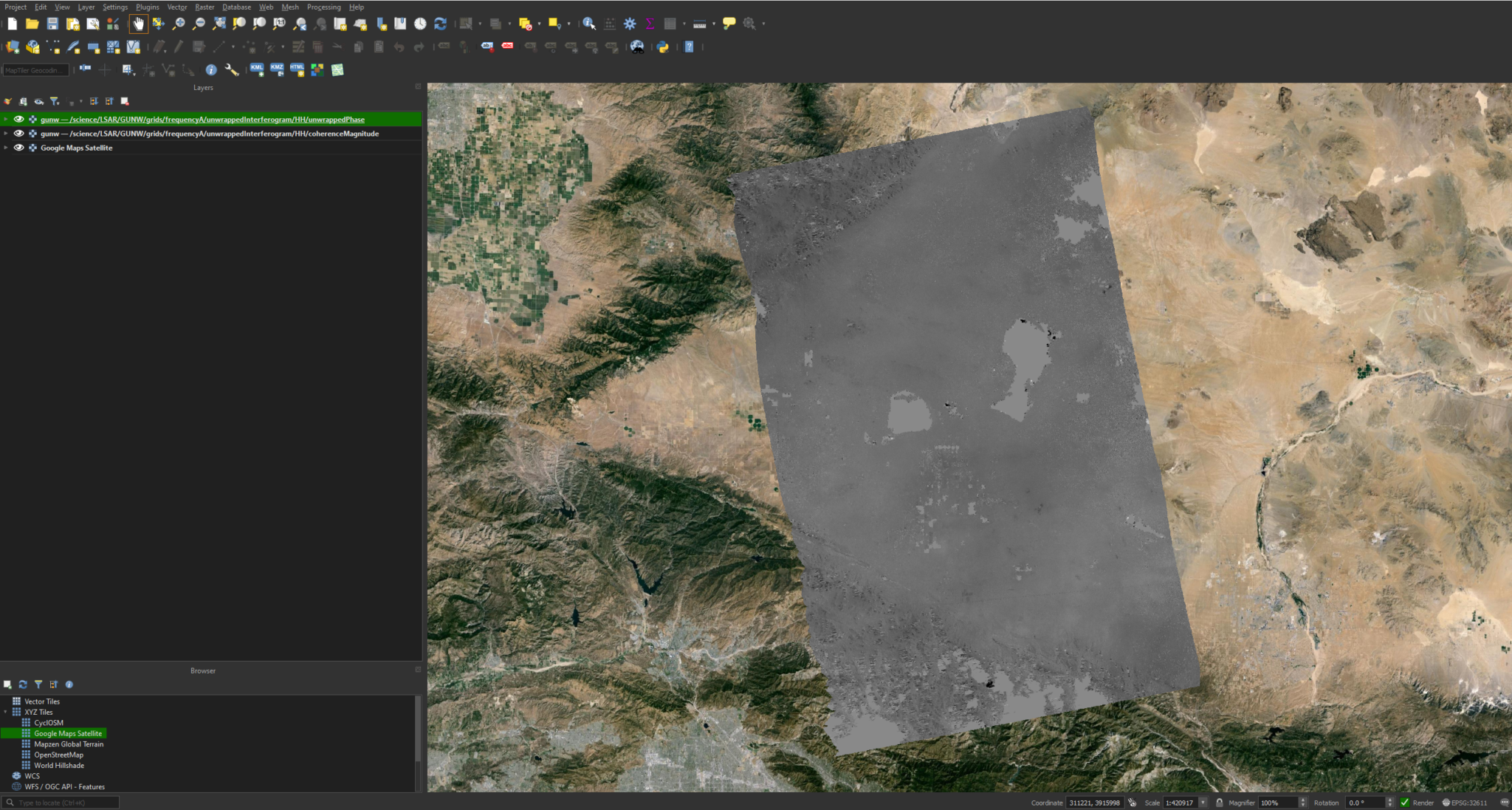
While this image is relatively clear and interpretable, we can modify the symbology to bring out some more detail. We can do this by right-clicking the layer and selecting Properties. We can then go to Symbology -> Min / Max Value Settings and select Cumulative Cut Count or Mean +/- standard deviation.

When color is desired, you can also switch the Render Type to singleband psuedocolor in the same menu.
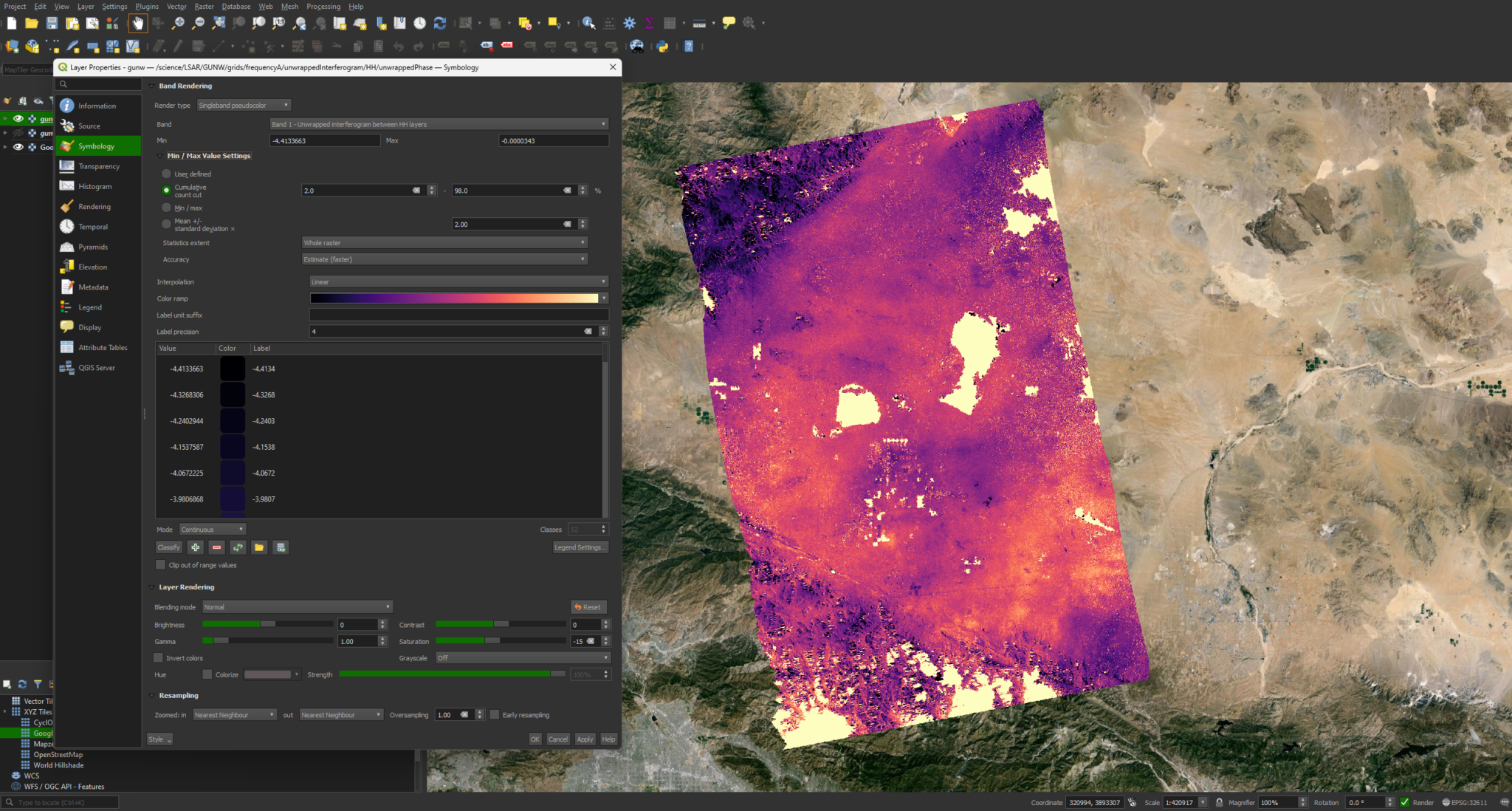
Coherence Magnitude
The coherence magnitude image should look good by default, but the same process that we applied to the unwrapped image can be applied if desired.

Wrapped Phase
Unlike the unwrapped phase and coherence magnitude, the wrapped phase data uses a complex data type. Unfortunately QGIS won’t natively handle complex data, so we will have to extract the phase component using this GDAL command:
gdal_translate -of GTiff DERIVED_SUBDATASET:PHASE:path/to/nisar.nc:/science/LSAR/GUNW/grids/frequencyA/wrappedInterferogram/HH/wrappedInterferogram phase.tif
We can then load the GeoTIFF image into QGIS as normal. It should look good by default, but you can modify the Symbology in the same manner as the unwrapped phase, if desired.

If you’re experiencing instant-crashes, even if you’ve changed the file extension to .nc, refer to the QGIS Crashes section.
The GOFF sample product is blank. Changing the file extension will allow the product to open correctly in QGIS, although there is no data in the images to demonstrate at the moment.
Some versions of QGIS using particular operating systems will crash when adding some of the NISAR sample products.
QGIS Version 3.36 on Windows is the only version tested that did not crash when either the GUNW or GOFF products were added to the project. With QGIS 3.36 on a Windows machine, you can load in the HDF5 files without renaming them to .nc.
On MacOS and Ubuntu, QGIS 3.36 will instantly crash when a NISAR dataset with HDF5 drivers is loaded. Earlier versions of QGIS also crash on Windows machines, as well as on MacOS and Ubuntu. This is due to the HDF5 versions being old/incorrect on those operating systems. It may be possible to fix this by downloading the new versions of the HDF5 drivers from HDF Group and replacing them. You can check if a dataset will have this issue by checking the driver using gdalinfo.
The error seems to be an invalid memory access (segfault) by QGIS. It appears that QGIS is continuing to load the images as HDF5, rather than NetCDF.
Panoply Exploration

This section demonstrates how to use the NASA Panoply software to explore NISAR sample data.
This documentation assumes you have installed Panoply, and downloaded the NISAR sample data to a directory named nisar_sample_data on your local machine. There are 4 Level-2 data files (GUNW, GOFF, GCOV, GSLC) and one Level-3 product (SME2). We are going to visualize each product.
Click on a product type to view the steps for Exploration of the sample data using Panoply:
The GUNW data file is
NISAR_L2_PR_GUNW_001_005_A_219_220_4020_SH_20081012T060910_20081012T060926_20081127T060959_20081127T061015_P01101_M_F_J_001.h5
1). Open the File
cd ${path_to_panoply}
./panoply.sh
The Panoply user interface opens as shown below.

- Navigate to the directory with the file in the Open window
- Click on the NISAR_L2_PR_GUNW_001_005_A_219_220_4020_SH_20081012T060910_20081012T060926_20081127T060959_20081127T061015_P01101_M_F_J_001.h5 file
- Click the
Openbutton to load the file into panoply as shown below
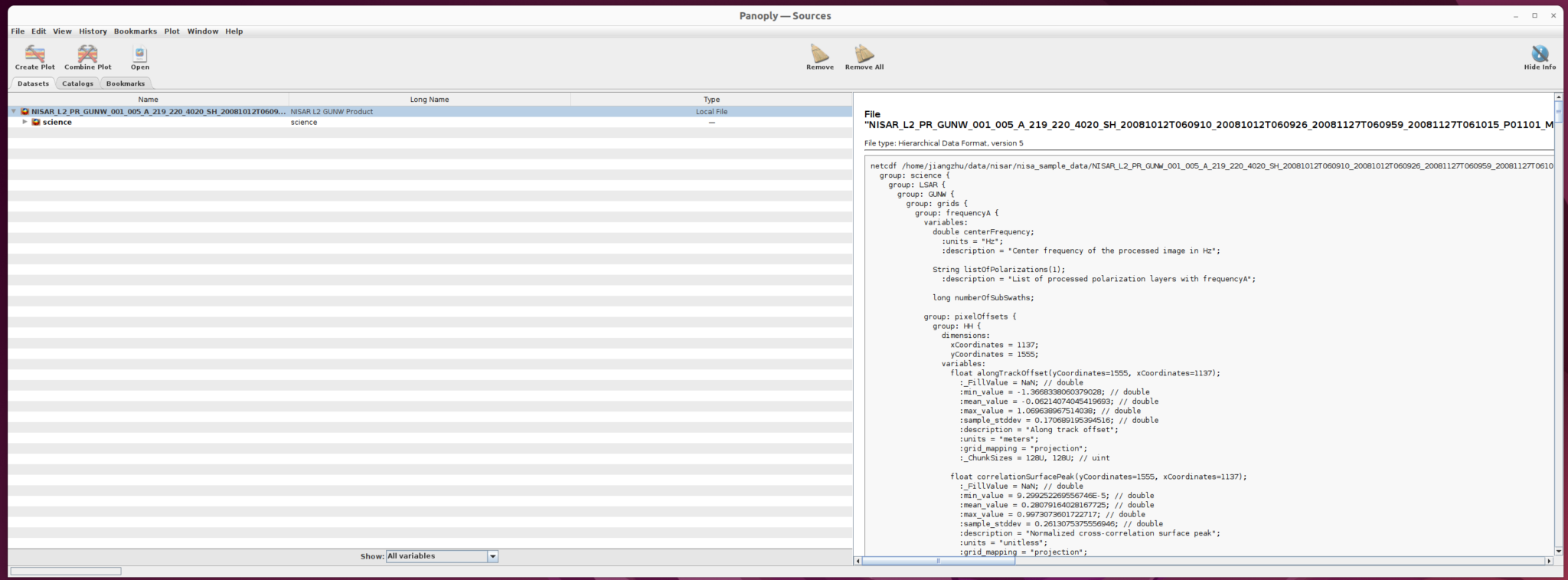
The right panel shows the structure of the file as well as the metadata. You can use this interface to inspect file information. The left panel provides users the option to select the dataset they want to use.
Expand the drop-down menu to find the “unwrappedPhase” dataset, which we will visualize in the following steps.

2) Create Plot
Once you select the dataset to use (in this case, unwrappedPhase), click the Create Plot button right below the main menu. A dialogue window opens with options for the type of plot to create.
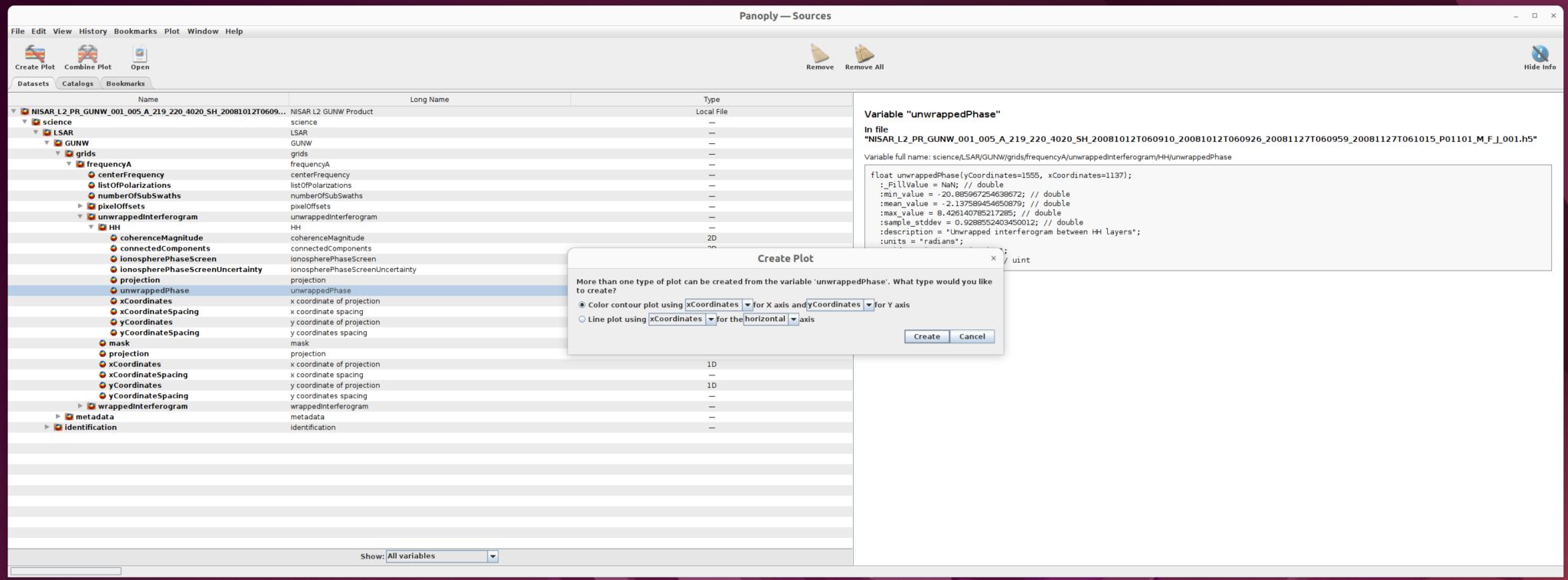
Select Color contour plot, then click the create button in the dialogue window to display the plot window.

3) Customize the Plot
Users can control the plot properties by clicking window in the pull-down menu of the plot window.
As an example, to change the scale to show off the variation of unwrappedPhase:
- Select
Plot Controlsto open the Plot Controls window. - Select
scale. - Change the min value to -8.426141, and keep the max 8.426141 unchanged.
The new plot shows off the variation clearly.
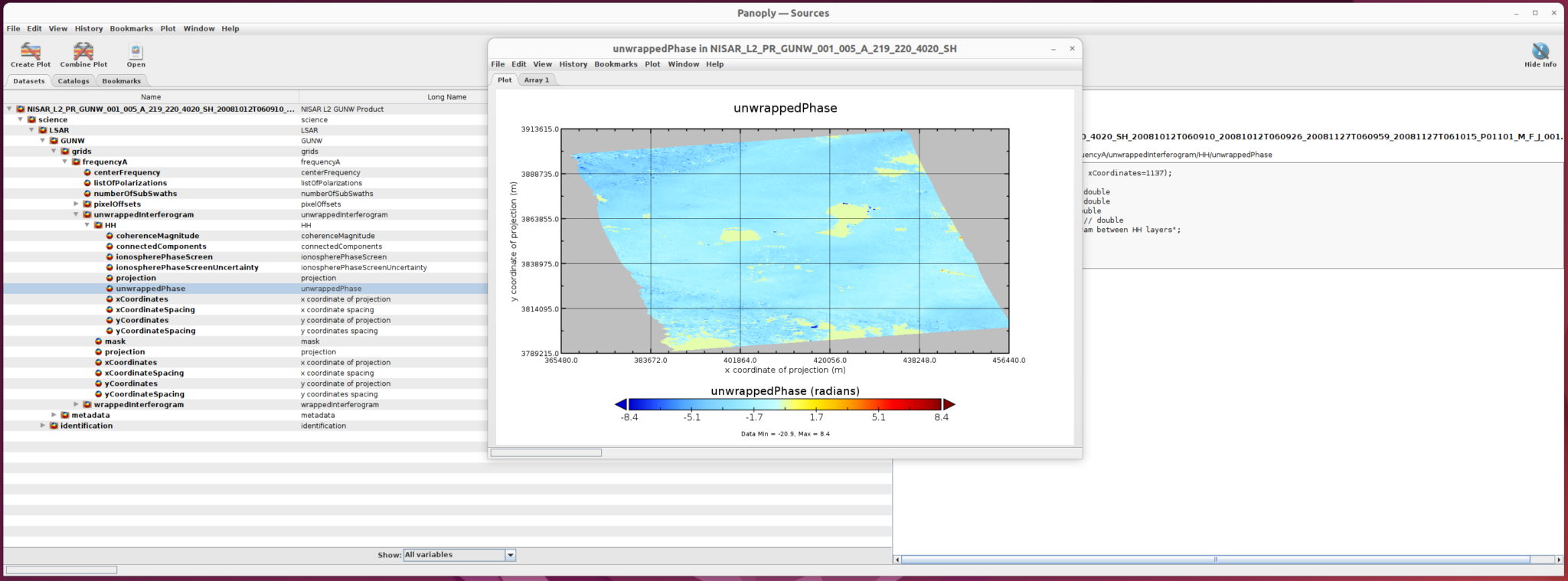
4) Save the Plot
When you are satisfied with the plot, you can save the results.
- Select
Filein the main menu of plot window - Select
Save Imageto open the Save As… window - Select the directory, and either accept the default filename or input the filename of your choice
- Click the
savebutton to save the plot

The example GOFF product is:
NISAR_L2_PR_GOFF_001_005_A_219_220_4020_SH_20081012T060910_20081012T060926_20081127T060959_20081127T061015_P01101_M_F_J_001.h5
Use the same process detailed in the GUNW Product section to visualize the GOFF product.

The example GCOV product is:
NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5
It is not possible to open the current example GCOV product directly in Panoply.
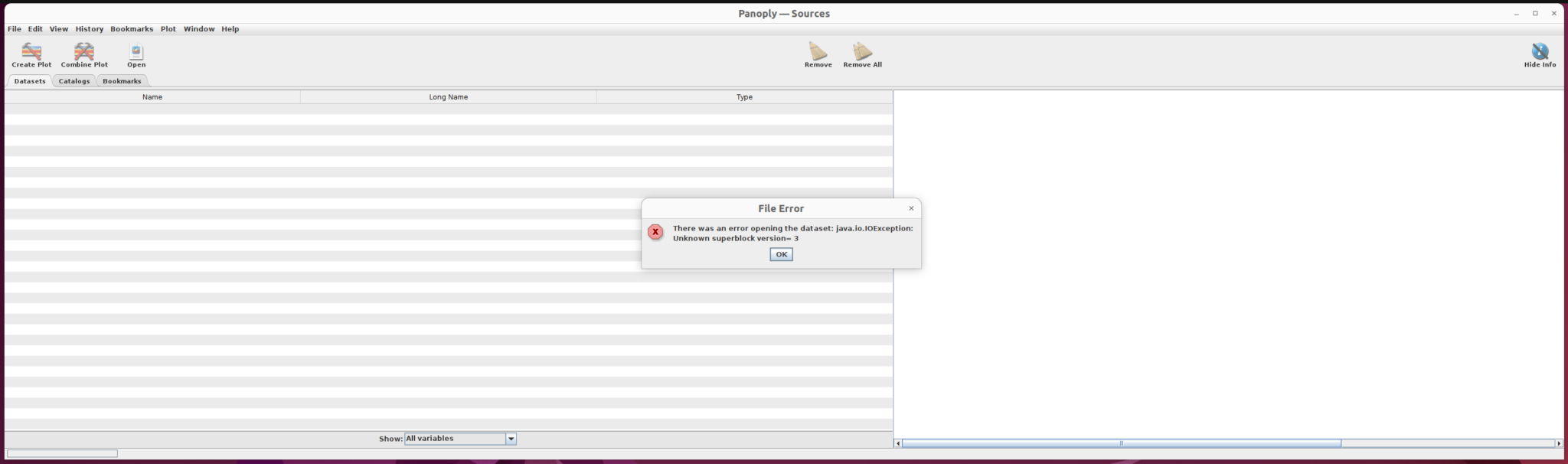
To visualize the GCOV data, use a GDAL command to extract the single backscatter subdataset from the h5 file and save it as a NetCDF file. Once the file type has been converted, we can use Panoply to visualize it.
1) Use gdalinfo to get the information about the individual sub-datasets
gdalinfo NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5
2) Use gdal_translate to convert the subdataset into a Netcdf file
gdal_translate -of NetCDF NETCDF: "NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5"://science/LSAR/GCOV/grids/frequencyA/HHHH HHHH.nc
gdal_translate -of NetCDF NETCDF:"NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5"://science/LSAR/GCOV/grids/frequencyA/rtcAreaNormalizationFactorGamma0ToSigma0 rtcAreaNormalizationFactorGamma0ToSigma0.nc
3) Load the NetCDF file in the Panoply to view the plot
HHHH.nc is displayed with a color contour plot using x for x and y for y

You may need to open the Plot Control window and adjust the grid, scale, and map projection in order to get suitable visualization.
The image below illustrates the adjustments made in the Plot Control window to visualize the rtcAreaNormalizationFactorGamma0ToSigma0.nc file.
rtcAreaNormalizationFactorGamma0ToSigma0.nc is displayed with color contour plot using x for x and y for y

If you choose to create Georeferenced longitude-latitude color contour plot. You may need to open the Plot Control window and adjust the grid, scale, and map projection in order to get suitable visualization.
The image below illustrates the adjustments made in the Plot Control window to visualize the rtcAreaNormalizationFactorGamma0ToSigma0.nc file.
This is the original display of rtcgcov_rtc_lonlat_countour.nc file in Panoply:

The video below illustrates the adjustments made in the Plot Control window to visualize the rtcgcov_rtc_lonlat_countour.nc file appropriately.
clip_gcov_subdataset_grid_scale_map.webm
After applying the adjustments, the rtcgcov_rtc_lonlat_countour.nc file is displayed as illustrated below.

The example GSLC product is:
NISAR_L2_PR_GSLC_001_005_A_219_2005_DHDH_A_20081127T060959_20081127T061015_P01101_F_N_J_001.h5
If we use the same process detailed in the GUNW Product section to visualize the GSLC product, we do not get useful results.
Instead of opening the GSLC product directly, we can extract the sub-datasets in the h5 file using GDAL, and then visualize these sub-datasets in Panolply.
The subdataset is a complex datatype, so we cannot use the gdal_translate command to convert a complex dataset directly into a NetCDF file. We use the following logic to generate a file that can be used to visualize the subdataset:
1) Convert the subdataset into a geotiff file
gdal_translate -of GTiff NETCDF:"NISAR_L2_PR_GSLC_001_005_A_219_2005_DHDH_A_20081127T060959_20081127T061015_P01101_F_N_J_001.h5"://science/LSAR/GSLC/grids/frequencyA/HH hh.tif
2) Convert the single-band Complex32 dataset into a two-band Float32 GeoTIFF file
from osgeo import gdal
import numpy as np
file="hh.tif"
outfile = "hh_polar.tif"
ds = gdal.Open(file)
band = ds.GetRasterBand(1)
data = band.ReadAsArray()
amp = abs(data)
phase = np.angle(data)
# create output raster file
driver = gdal.GetDriverByName("GTiff")
outdata = driver.Create(outfile, ds.RasterXSize, ds.RasterYSize, 2, gdal.GDT_Float64)
outdata.SetGeoTransform(ds.GetGeoTransform())
outdata.SetProjection(ds.GetProjection())
outdata.SetMetadata(ds.GetMetadata())
outdata.GetRasterBand(1).WriteArray(amp)
outdata.GetRasterBand(1).SetMetadata({"bandname": "modulus"})
outdata.GetRasterBand(2).WriteArray(phase)
outdata.GetRasterBand(2).SetMetadata({"bandname": "argument"})
outdata = None
3) Convert the GeoTIFF file to a NetCDF file
gdal_translate -of NetCDF hh_polar.tif hh_polar.nc
4) Use Panoply to visualize the NetCDF file

The SME2 product sub-datasets can be displayed using the same process that we used for the GUNW product.
For example, the soil moisture sub-dataset is displayed here:

and the landcover sub-dataset is displayed here:
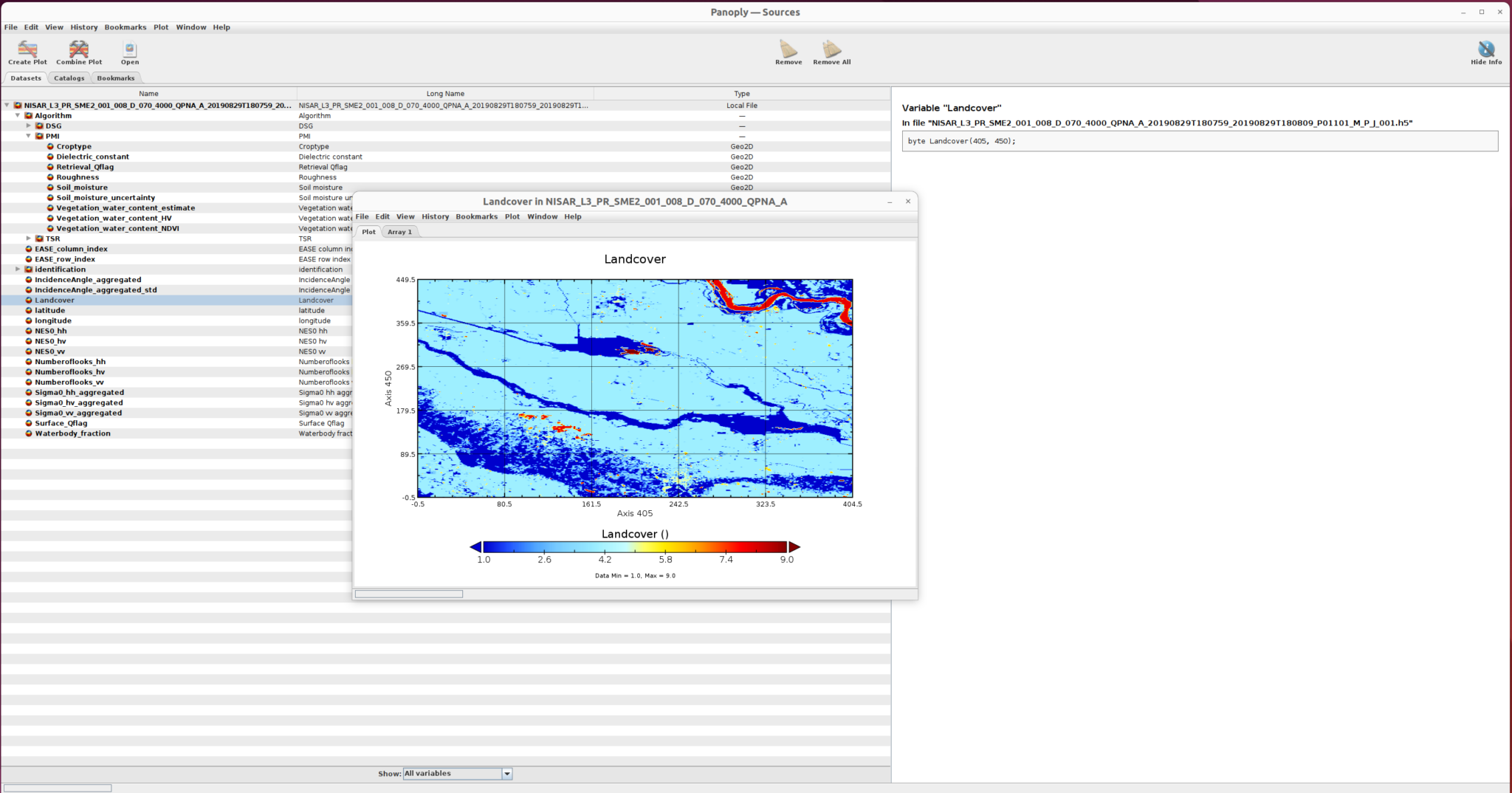
GDAL Data Access/Transfomation

This section guides accessing and transforming NISAR HDF5-formatted data with GDAL, using the GCOV sample dataset as the primary test case.
One important thing to remember is that the NISAR HDF5 products are actually more similar to NetCDFs. Notably, the spatial reference data is encoded using NetCDF Climate and Forecast (CF) Metadata Conventions. This means you need to specify an output format for some commands, otherwise GDAL will throw an error like ERROR 1: Cannot guess driver for output.h5. It also means that you should prepend the NISAR files with NETCDF:. This approach is demonstrated throughout the examples in this section. Note that while the formats are slightly different, you can still access the sub-datasets or groups the same way you would with an HDF5 file.
For more detailed info on what each of the products contain, and for sample data, visit https://nisar.jpl.nasa.gov/data/sample-data/.
Run this command to get a list of the sub-datasets and the paths that you need to use with your commands:
gdalinfo path/to/NISAR.h5 | grep "SUBDATASET_\d*_DESC"
Converting the NISAR data will require you to specify a sub-dataset (also known as a group). Failure to supply one will result in an error like Input file contains subdatasets. Please, select one of them for reading.
The examples below choose to convert only the HH to HH covariance portion of GCOV product. This data is most similar to our current co-polarizated RTC products.
Using the Command Line
gdal_tranlsate -of GTiff NETCDF:"path/to/input.h5":/science/LSAR/GCOV/grids/frequencyA/HHHH path/to/output.tif
Using Python
h5_file = 'NETCDF:"path/to/input.h5":/science/LSAR/GCOV/grids/frequencyA/HHHH'
out_file = 'path/to/output.tif'
gdal.Translate(out_file, h5_file, format='GTiff')
Reprojecting the NISAR data will require you to specify which sub-dataset you would like to reproject. Failure to specify the sub-dataset will result in an error like ERROR 1: Input file NETCDF:NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5 has no raster bands.
Using the Command Line
gdalwarp -of GTiff -t_srs EPSG:XXXX NETCDF:"path/to/input.h5":/science/LSAR/GCOV/grids/frequencyA/HHHH path/to/output.tif
Using Python
h5_file = 'NETCDF:"path/to/input.h5":/science/LSAR/GCOV/grids/frequencyA/HHHH'
out_file = 'path/to/output.tif'
gdal.Warp(tif_file, h5_file, format='GTiff', dstSRS='EPSG:XXXX')At this time, we do not know of a simple way of reprojecting an entire dataset. You will need to individually reproject each subdataset that you are interested in.
A sub-dataset can be clipped to a geographic extent using gdalwarp. The existing exent of the sub-dataset can be found by calling gdalinfo on it. Note that you won’t find the extent by calling gdalinfo on the entire dataset.
Using the Command Line
gdalwarp -of GTiff -te xmin ymin xmax ymax NETCDF:"path/to/input.h5":/science/LSAR/GCOV/grids/frequencyA/HHHH path/to/output.tif
Using Python
h5_file = 'NETCDF:"path/to/input.h5":/science/LSAR/GCOV/grids/frequencyA/HHHH'
out_file = 'path/to/output.tif'
gdal.Warp(out_file, h5_file, format='GTiff', outputBounds=[xmin, ymin, xmax, ymax])Note that the Command Line method only adds the main images to the multiband GTiff without the H5s additional metadata. The Python method is more complicated, but it will place all subdatasets into the GTiff, as well as the additional related metadata.
Using the Command Line
file="path/to/input.h5"
subdataset1="${file}://science/LSAR/GCOV/grids/frequencyA/HHHH"
subdataset2="${file}://science/LSAR/GCOV/grids/frequencyA/numberOfLooks"
subdataset3="${file}://science/LSAR/GCOV/grids/frequencyA/rtcAreaNormalizationFactorGamma0ToSigma0"
dimensions_args="-ul_lr upper_left_corner_x upper_left_corner_y lower_right_corner_x lower_right_corner_y -ps pixel_size_x pixel_size_y"
gdal_merge.py $dimensions_args -seperate -of GTiff -o $(echo $file | sed 's/\.h5/\_multiband.tif/g')
Using Python
Here is the code to extract raster subsets in h5 and convert them into geotiff files.
import json
import subprocess
import re
from osgeo import gdal
file="NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5"
actual_json = json.loads(subprocess.check_output(f"gdalinfo -json {file}", shell=True))
datasets = actual_json['metadata']['SUBDATASETS']
names = [key for key in datasets.keys() if re.match('SUBDATASET_[0-9]*_NAME', key)]
for name in names:
subdata = f"{datasets[name].replace('HDF5','NETCDF')}"
if gdal.Open(subdata).GetProjectionRef() != '':
outfile =f"{subdata.replace('/','_')}.tif"
subprocess.run(f'gdal_translate -of GTiff {subdata} {outfile}', shell=True)
We also provide the code to combine all raster subdatasets with the same datatype, extent, size, and projection in the h5 into multiple-band geotiff files. We notice that the combined multiple band geotiff files do not include the metadata for each band in the geotiff files.
import json
import subprocess
import re
import numpy as np
from osgeo import gdal
def combine_geotiff(filelist, outfile):
'''
Merge the geotiff files with the same data type, extent, and projection into a multiple band geotif file
:param filelist: list of geotiff files
:param outfile: filename of the output file
:return: None
'''
bandmetalist = [gdal.Info(file, format='json')['bands'] for file in filelist]
bandnumlist = [len(item) for item in bandmetalist]
bandnum = int(np.array(bandnumlist).sum())
filenum = len(filelist)
for i in range(filenum):
ds = gdal.Open(filelist[i])
if i == 0:
# create output raster file
driver = gdal.GetDriverByName("GTiff")
outdata = driver.Create(outfile, ds.RasterXSize, ds.RasterYSize, bandnum, ds.GetRasterBand(1).DataType )
outdata.SetGeoTransform(ds.GetGeoTransform()) ##sets same geotransform as input
outdata.SetProjection(ds.GetProjection()) ##sets same projection as input
bandnum_of_file = ds.RasterCount
fmeta = ds.GetMetadata()
for j in range(1, bandnum_of_file + 1):
band = ds.GetRasterBand(j)
outdata.GetRasterBand(i*bandnum_of_file + j).WriteArray(band.ReadAsArray())
fmeta.update(band.GetMetadata())
outdata.GetRasterBand(i*bandnum_of_file + j).SetMetadata(fmeta)
ds = None
outdata = None
file="NISAR_L2_PR_GCOV_001_005_A_219_4020_SHNA_A_20081012T060910_20081012T060926_P01101_F_N_J_001.h5"
actual_json = json.loads(subprocess.check_output(f"gdalinfo -json {file}", shell=True))
datasets = actual_json['metadata']['SUBDATASETS']
names = [key for key in datasets.keys() if re.match('SUBDATASET_[0-9]*_NAME', key)]
datalist=[]
for name in names:
subdata = f"{datasets[name].replace('HDF5','NETCDF')}"
ds = gdal.Open(subdata)
if ds.GetProjectionRef() != '':
size = f'{ds.RasterXSize}x{ds.RasterYSize}'
outfile = f"{subdata.replace('/','_')}.tif"
outfile = outfile.split(":")[2].replace('__','')
mo=outfile.replace(".tif","")
subprocess.run(f"gdal_translate -of GTiff -ot Float32 -mo Band={mo} {subdata} {outfile}", shell=True)
datalist.append({"filename":outfile, "size":size})
sizelist = [item['size'] for item in datalist]
uniqvals= np.unique(sizelist)
for item in uniqvals:
itemlist = [ dataitem['filename'] for dataitem in datalist if dataitem['size'] == item ]
mb_outfile = subdata.split(":")[1].replace('.h5',f'_{item}.tif')
mb_outfile = mb_outfile.replace('"','')
combine_geotiff(itemlist, mb_outfile)